Modeling Count Data with Poisson Regression¶
How to do the regression?¶
We model the count data as
How to optimize?¶
We need to find the best \(\beta\), thus we maximize the likelihood function. The count data can be generalized with the Poisson distribution, hence
It's hard to optimize the \(\prod\) operation, thus we \(\ln\) it.
Our objective is to minimize the negative log likelihood, \(\text{nll}(\beta)\) Taking the derivative of \(\text{nll}(\beta)\) with respect to \(\beta\) we get
We need to optimize Eq. (3) iteratively with Gradient Descent
Build Functions¶
Function to predict the count¶
We model the count data as
- We do the vectorized operations of \(X\) and \(\beta\). This functions will return an array with shape of
(n_samples,)
Function to calculate the negative log likelihood¶
We calculate the model performance by the negative log likelihood
- There is no broadcasting for factorial function thus we use
np.math.factorial(val)and list comprehension to calculate \(\ln (y_{i}!)\) - We know that \(\hat{y}_{i} = \exp(X_{i} \beta)\), then \(X_{i} \beta = \ln (\hat{y}_{i})\)
Modeling Example - Horseshoe Crab¶
We want to model the count of Horseshoe crab satellites. We get the data from here.
Load the data¶
Save the data to an excel format, then load the data
Data shape: (173, 5)
Color Spine Width Weight Satellite
0 2 3 28.3 3.05 8
1 3 3 26.0 2.60 4
2 3 3 25.6 2.15 0
3 4 2 21.0 1.85 0
4 2 3 29.0 3.00 1
There are 173 number of samples with 5 columns.
What we want to predict¶
We predict the count of Satellite in Horseshoe crabs.
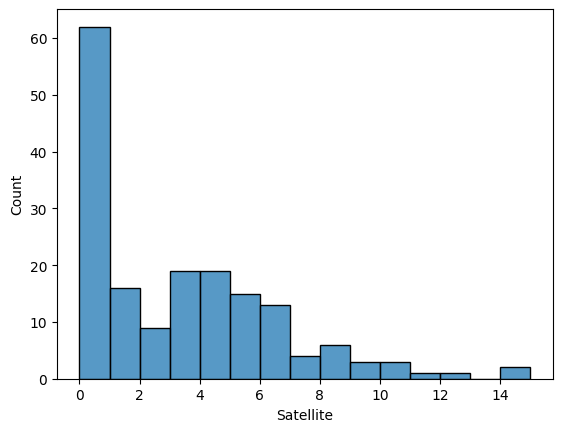
Split input & output data¶
First, create a function to split input & output
- For simplicity, we only choose one prediction, i.e.
Width
Print the splitting results
Prepare for the optimization¶
We want to optimize \(\beta\). Recall the Equation (1)
In a vectorized form, we have
We have to recreate \(\mathbf{A}\) first,
[[ 1. 28.3]
[ 1. 26. ]
[ 1. 25.6]
[ 1. 21. ]
[ 1. 29. ]
[ 1. 25. ]
[ 1. 26.2]
[ 1. 24.9]
[ 1. 25.7]
[ 1. 27.5]]
We will use \(\mathbf{A}\) and \(\mathbf{y}\) during the optimization process.
Next, we have to initialize the model parameters \(\beta\)
The Gradient Descent¶
Our objective is to minimize the negative log likelihood, hence we update the \(\beta\):
We do this iteratively, thus
-
This is a stopping criterion. When
i == max_iter-1the optimization process will stop. -
learning_ratewill control how big the changes is when updating the model parameter during optimization process. A big value oflearning_ratecan make the model parameter updates diverges. -
Predict the count using Eq. (1)
-
Predict the negative log likelihood using Eq. (2)
-
Predict the gradient of NLL using Eq. (3). We use vectorized operation, i.e.
np.dot(A, B), instead ofnp.sum(A * B). -
Update the model parameter using Eq. (4)
-
Print the optimization process. You can monitor the NLL. There's something wrong if your NLL is getting bigger.
iter:1, nll: 2013.5065, beta:[-2.9166 0.0671]
iter:10001, nll: 461.7734, beta:[-2.9746 0.1519]
iter:20001, nll: 461.7201, beta:[-3.0261 0.1538]
iter:30001, nll: 461.6821, beta:[-3.0696 0.1554]
iter:40001, nll: 461.6551, beta:[-3.1063 0.1568]
iter:50001, nll: 461.6358, beta:[-3.1373 0.1579]
...
iter:450001, nll: 461.5881, beta:[-3.3046 0.164 ]
iter:460001, nll: 461.5881, beta:[-3.3046 0.164 ]
iter:470001, nll: 461.5881, beta:[-3.3046 0.164 ]
iter:480001, nll: 461.5881, beta:[-3.3046 0.164 ]
iter:490001, nll: 461.5881, beta:[-3.3047 0.164 ]
Finally, we get our trained model parameter \(\hat{\beta}\)
Evaluation¶
Let's check its performance. First, we make a prediction
[3.81032936 2.61279026 2.44685093 1.15051312 4.2739785 2.21749145
2.6999335 2.18141199 2.48732062 3.34170756 2.65600451 4.20443917
...
3.62735231 3.87335031 8.9417806 2.52845967 1.88199888 1.62368218
3.81032936 2.83612812 2.83612812 2.65600451 2.04286965] # (1)
- The results of Poisson model is not
int
We cannot compare anything right now, let's plot the count distribution of Satellite between actual & predicted value.
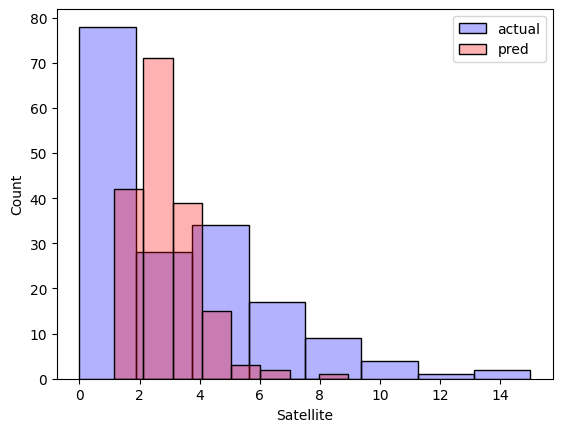
Not a good prediction though.
Modeling with Statsmodel¶
Now, let's try modeling with a well-known library: statsmodels
Do the modeling
- Poisson regression is a family of GLM (generalized linear model).
Then print the results
Generalized Linear Model Regression Results
==============================================================================
Dep. Variable: Satellite No. Observations: 173
Model: GLM Df Residuals: 171
Model Family: Poisson Df Model: 1
Link Function: Log Scale: 1.0000
Method: IRLS Log-Likelihood: -461.59
Date: Sat, 17 Jun 2023 Deviance: 567.88
Time: 14:28:46 Pearson chi2: 544.
No. Iterations: 5 Pseudo R-squ. (CS): 0.3129
Covariance Type: nonrobust
==============================================================================
coef std err z P>|z| [0.025 0.975]
------------------------------------------------------------------------------
const -3.3048 0.542 -6.095 0.000 -4.368 -2.242
x1 0.1640 0.020 8.216 0.000 0.125 0.203
==============================================================================
As we can see, the coefficient is [-3.3048, 0.1640] similar to our from scratch results.
Eventhough the results is not good enough, however we can validate that our from scratch works fine compared to the statsmodels library.